Doug Barrick (he)
Thomas C. Jenkins Professor in Biophysics and Chair
Contact Information
- barrick@jhu.edu
- 216 Jenkins Hall
- 410-516-0409
- Group/Lab Website
Research Interests: Statistical thermodynamics in biological systems; stability and folding of modular proteins, relations between amino acid sequence, consensus, and evolution; physical chemistry, structure, and function in Notch signaling.
Education: PhD, Stanford University
Doug Barrick is Professor and Chair of the Jenkins Department of Biophysics. He has been on the faculty in the Department of Biophysics since 1997. He obtained his BA at the University of Colorado, studying Molecular Biology and Chemistry, where he did independent research with Larry Gold, Gary Stormo, and Tom Schneider, and learned information theory, multiple regression techniques, nearest-neighbor models, and recombinant DNA work.
Dr. Barrick obtained his Ph.D. in the Department of Biochemistry at Stanford in the laboratory of Buzz Baldwin, where he studied protein folding, and experimentally and thermodynamically characterized a partly folded “molten globule” state of apomyoglobin. He did postdoctoral research at the University of Oregon in the laboratory of Rick Dahlquist, where he used chemical rescue techniques to study structure, energetics, and allostery in myoglobin and hemoglobin, and learned some NMR.
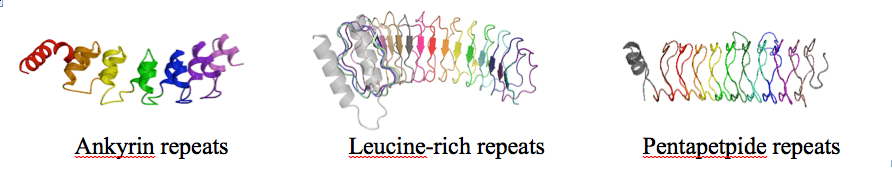
At Johns Hopkins University, Dr. Barrick and his lab have been focusing on the folding of repeat proteins, and on the structure, energetics, and function of Notch signaling. The folding studies have involved a series of roughly translationally symmetric proteins that can be fragmented and extended, allowing direct correlations to be made between stability, folding kinetics, and protein structure. Using variable-length proteins made of identical “consensus”-based repeats, the Barrick lab has been able to apply nearest-neighbor “Ising” models to resolve intrinsic stability from coupling between repeats. This approach is providing a direct experimental quantification of cooperativity in protein folding, an important but difficult-to-characterize quantity.
These studies have also given rise to a more detailed study of consensus stabilization in globular proteins. The Barrick lab has recently had some surprising successes with this approach, and is currently extending and refining methods for consensus sequence design, to better understand why it works, why it sometimes fails, and how to make it work better. With these insights, we hope to be able to design high-value pharmaceutical and industrial proteins with improved stabilities, solubilities, and shelf-life.
The Barrick lab has also had a long-standing project to understand the complex network of interactions that make up the Notch signaling pathway. Notch signaling is a key pathway in metazoan (including human) cell fate determination. Defects in Notch signaling have been implicated in a wide variety of diseases, most importantly, in various forms of cancer. We have been using a combination of structural and biophysical techniques, including x-ray crystallography, NMR spectroscopy, analytical ultracentrifugation, ITC, cell-culture transcription and ubiquitination assays, and most recently, single-molecule fluorescence spectroscopy to learn how the many conformational transitions, including disorder-order transitions and bivalent binding, lead to a precisely controlled signal response.
Outside the lab, Dr. Barrick has written a textbook entitled “Biothermodynamics: from Theory to Application”, which was published in 2017. As with his teaching, this text focuses on using software and physical examples to illustrate complex problems in physical chemistry in an intuitive way, and to give students a hands-on approach to this analysis. Another area of emphasis is to directly analyze data using various fitting procedures to explore models and measure key chemical and biological properties.
Dr. Barrick has received numerous research awards and grants, including graduation Summa Cum Laude, a HHMI Predoctoral Fellowship, a Helen Hay Whitney Postdoctoral Fellowship, an Arnold and Mabel Beckman Young Investigator Award, and a Johns Hopkins Teaching Award. Doug has served as a charter member of two NIH study sections (ICP1 and MSFB) and served as Chairperson for both sections. Doug currently serves as co-chair of the Scientific Advisory Committee of the Arnold and Mabel Beckman Foundation.
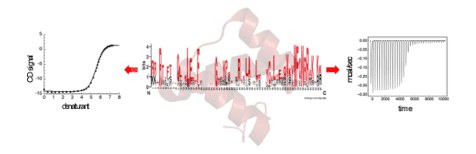
Dr. Barrick and his lab have been focusing on the folding of repeat proteins, and on the structure, energetics, and function of Notch signaling. The repeat-protein folding studies,funded by the NIH (see grant 1R01 GM068462), have involved a series of roughly translationally symmetric proteins that can be fragmented and extended (see below), allowing direct correlations to be made between stability, folding kinetics, and protein structure.
Using variable-length proteins made of identical “consensus”-based repeats, the Barrick lab has been able to apply nearest-neighbor “Ising” models to resolve intrinsic stability from coupling between repeats. This approach is providing a direct experimental quantification of cooperativity in protein folding, an important but difficult-to-characterize quantity.
These studies have also given rise to a more detailed study of consensus stabilization in globular proteins. The Barrick lab has recently had some surprising successes with this approach. Our first success is a consensus homeodomain that is super-stable and binds DNA with 100-fold increased affinity (currently under review at JACS).
We are currently extending methods for consensus sequence design, to better understand why consensus design works, why it sometimes fails, and how improve it. With these insights, we hope to be able to design high-value pharmaceutical and industrial proteins with improved stabilities, solubilities, and shelf-life, and to improve our understanding of the relationships between protein sequence and evolution, stability, dynamics, allostery, and function. Doug has submitted an NSF grant to support this research.
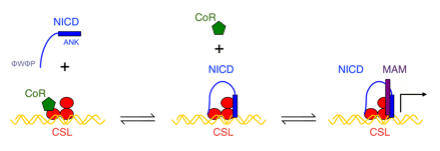
The Barrick lab has also had a long-standing project to understand the complex network of interactions that make up the Notch signaling pathway, funded by the NIH (see grant 1R01 GM060001). Notch signaling is a key pathway (see figure below) in human cell fate determination. Notch defects have been implicated in a variety of diseases, including cancer. We use a combination of structural and biophysical techniques, including x-ray crystallography, NMR spectroscopy, analytical ultracentrifugation, ITC, cell-culture transcription and ubiquitination assays, and single-molecule spectroscopy to learn how the many conformational transitions, including disorder-order transitions and bivalent binding, lead to a precisely controlled signal response.
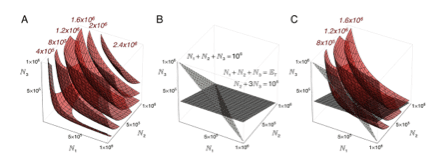
Outside the lab, Doug has put significant effort into teaching, and is nearly finished writing a text entitled “Biophysical Chemistry: a Visual Approach". As with his teaching, this text focuses on using software and physical examples to illustrate complex problems in physical chemistry in an intuitive way, and to give students a hands-on approach to this analysis. The figure below shows a visual representation of understanding how the Boltzmann distribution (C) results from maximizing multiplicity (A) subject to constraints on ensemble size and energy (B).
Another area of emphasis is to directly analyze data using various fitting procedures to explore models and measure key chemical and biological properties.
Doug teaches various aspects of biophysical chemistry to undergraduates and graduate students. His main teaching effort is a semester long course in undergraduate physical chemistry, using macromolecular examples. This course emphasizes hands-on learning; in addition to three weekly lectures, the course includes includes an hour-long weekly, computer-oriented small group recitation led by Doug. Students learn to do math, analyze equations, plot 3D functions, and analyze data using nonlinear least squares methods, as well as roll dice, collect data, and play statistical thermodynamics-based card games. This course is currently taught to upper-level undergraduates in the Biophysics and Chemistry departments, entitled Biophysical Chemistry (AS.250.372) and Physical Chemistry I with Biophysical Applications (AS.030.370).
Doug participates several graduate student courses. These include lectures on chemical kinetics in Physical Chemistry of Biological Macromolecules (AS.250.689), lectures on spectroscopy in Methods in Molecular Biophysics (AS.250.690). In addition, Doug is giving a one week "bootcamp" style module in the spring entitled Statistics and Data analysis (AS.250.622), and participates in the modules Solution Biophysics (AS.250.620) and Introduction to Computing in Biology (AS.250.649)
Full publication list available on Barrick Lab website.
- Geiger-Schuller, K.P., Mitra, J., Ha, T., & Barrick, D (2019) Functional instability allows access to DNA in longer Transcription Activator-Like Effector (TALE) arrays. eLife 8. pii: e38298.
- Geiger-Schuller, K., Sforza, K., Yuhas, M., Parmeggiani, F., Baker, D., & Barrick, D. (2018) Extreme stability in de novo-designed repeat arrays is determined by unusually stable short-range interactions. Proc. of the Natl. Acad. of Sci. USA 115, 7539-7544.
- Sherry, K.P., Das, R., Pappu, R.V., & Barrick D (2017) Control of transcriptional activity by design of charge patterning in the intrinsically disordered RAM region of the Notch Receptor. Proc. Nat. Acad. Sci. USA 114, E9243-E9252.
- Tripp, K. W., Sternke, M., & Barrick, D. (2017) Creating a homeodomain with high stability and DNA binding by sequence averaging. J. Am. Chem. Soc., 139, 5051-5060.
- Geiger-Schuller, K., & Barrick (2016) Broken TALES: Transcriptional activation-like effectors populate partly folded states. Biophysical Journal 111, 2395-2403.
- Fossat, M.J., Dao, T.P., Kenkins, K., Dellarole, M., Yang, Y., McCallum, S.A., Garcia, A.E., Barrick, D., Roumestand, C., & Royer, C.A, (2016) High-resolution mapping of a repeat protein folding free energy landscape, Biophysical Journal 111, 2368-2376 .
- Cunha, E., Hatem, C.L, & Barrick, D. (2016) Synergistic enhancement of cellulose pairs linked by consensus ankyrin repeats: determination of the roles of spacing, orientation, and enzyme identity. Proteins: Struct. Funct.& Bioinf. 84, 1043-1054.
- Marold, J.D., Kavran, J.M., Bowman, G.D., & Barrick, D. (2015). A naturally occurring repeat protein with high internal sequence identity defines a new class of TPR-like proteins. Structure 23, 2206-2265.
- Sherry, K.P., Johnson, S.E., Hatem, C.L., Majumdar, A., Johnson, S., Hatem, D. & Barrick, D. (2015). Effects of linker length and transient secondary structure elements in the intrinsically disordered RAM region on Notch signaling. J. Mol. Biol. 427, 3587-3597.
- Dao, T., Majumdar, A., & Barrick D. (2015) The highly polarized C-terminal transition state of the leucine-rich repeat domain of PP32 is governed by local stability. PNAS, 112 (18), E2298-306.
- Preimesberger, M.R., Majumdar, A., Aksel, T., Sforza, K., Leckta, T., Barrick, D, & Lecomte, J.T.J. (2015) Direct NMR Detection of Bifurcated Hydrogen Bonding in the a-Helix N-Caps of Ankyrin Repeat Proteins. J. Am. Chem. Soc., in press.
- Aksel, T., & Barrick, D (2014) Direct observation of parallel folding pathways revealed using a symmetric repeat protein system. Biophysical Journal 107, 220-232.
- Dao, T., Majumdar, A., & Barrick, D (2014). The role of the capping motifs in stabilizing the structure of the PP32 leucine-rich repeat domain. Protein Science 23 (6), 801-811.
- Cunha, E., Hatem, C., & Barrick, D (2013) Insertion of Endocellulase Catalytic Domains into Thermostable Consensus Ankyrin Scaffolds: Effects on Stability and Cellulolytic Activity. Applied Environmental Microbiology 79, 6684-6696.
- Johnson, S.E. & Barrick, D. (2012) Dissecting and cirumventing the requirement for RAM in Notch signaling. PLos One 7(8) 39093.
- Allgood, A.G, & Barrick, D. (2011) Mapping the Deltex binding surface on the Notch Ankyrin domain using analytical ultracentrifugation. J. Mol. Biol. 414, 243-259.
- Cunha, E., Hatem, C.L., & Barrick, D. (2011). Natural and designed enzymes for cellulose degradation, in Advanced Biofuels and Bioproducts, Springer Publishing, 339-370.
- Vieux, E.F., & Barrick, D. (2011). Deletion of internal structured repeats increases the stability of a leucine-rich repeat protein, YopM. Biophysical Chemistry 159, 152-161.
- Rouget, J. B., Aksel, T., Roche, J., Saldana, J. L., Garcia, A. E., Barrick, D., & Royer, C. A. (2011). Size and Sequence and the volume change of protein folding. J. Am. Chem. Soc., 133, 6020-6027.
- Sosnick, T.R., & Barrick, D. (2011). The folding of single domain proteins – have we reached a consensus? Curr. Op. Struct. Biol. 21, 12-24.
- Aksel, T., Majumdar, A., & Barrick, D. (2011) The contribution of entropy, enthalpy, and hydrophobic desolvation ito cooperativity in repeat-protein folding. Structure, 19, 349-360.
- Johnson, S.E. Ilagan, M.X.G., Kopan, R., & Barrick, D (2010) Thermodynamic analysis of the CSL:Notch interaction: Distribution of binding energy of the Notch RAM region to the CSL beta-trefoil domain and the mode of competition with the viral transactivator EBNA2. J. Biol. Chem, 285, 6681-6692.
- Kloss, E. & Barick, D. (2009) C-terminal deletion of leucine-rich repeats from YopM reveals a heterogeneous distribution of stability in a cooperatively folded protein. Protein Science, 18, 1948-1960.
- Aksel, T., & Barrick, D. (2009) Analysis of repeat-protein folding using nearest-neighbor statistical mechanical models. Meth. Enzymol. 455, 95-125.
- Street, T.O., Bradley, C.M., & Barrick, D. (2009) Predicting repeat-protein folding kinetics from an experimentally determined folding energy landscape. Prot. Sci. 18, 58-68.
- Kloss, E., & Barrick, D. (2008) Thermodynamics, kinetics, and salt dependence of folding of YopM, a large leucine-rich repeat protein. J. Mol. Biol. 383, 1195-1209.
- Courtemanche, N., & Barrick, D. (2008) The leucine-rich repeat domain of Internalin B folds along a polarized N-terminal pathway. Structure 16, 705-712.
- Tripp, K.W., & Barrick, D. (2008) Rerouting the folding pathway of the Notch ankyrin domain by reshaping the energy landscape. J. Am. Chem. Soc. 130, 5681-5688.
- Bertagna, A., Toptygin, D., Brand, L., & Barrick, D. (2008) The effects of conformational heterogeneity on the binding of the Notch intracellular domain to effector proteins: a case of biologically tuned disorder. Biochem. Soc. Trans. 36, 157-166.
- Courtemanche, N., & Barrick, D. (2008) Folding thermodynamics and kinetics of the Leucine-rich repeat domain of the virulence factor Internalin B. Protein Science 17, 43-53.
- Kloss, E., Courtemanche, N., & Barrick, D. (2007) Repeat-protein folding: new insights into origins of cooperativity, stability, and topology. Archives of Biochemistry and Biophyics 469, 83-99.
- Street, T.O., Courtemanche, N., & Barrick, D. (2007B) Protein folding and stability using denaturants. Methods in Cell Biology: Biophysical Tools for Biologists. Correia, J.J. & Detrich, H.W. III, eds. Academic Press, Vol 84, 295-325.